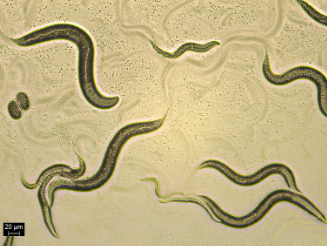
This blogpost is a guest contribution from researchers at European centre of excellence for sustainable water technology (Wetsus) in the Netherlands. In this post, Antoine Karengera talks about their recently published paper dealing with the potential use of the nematode – Caenorhabditis elegans – as a test organism for detecting toxic potencies of pollutants in water sources.
The problem
There are many contaminants in water sources that cannot all be efficiently measured by current chemical methods. Hydrophilic compounds are especially hard to extract, and unknown compounds, metabolites and reaction products are difficult to identify. Also, analytical techniques cannot provide information about the potential toxicity of the compounds and mixtures thereof. Although bioanalytical tools can quantify the toxic potency of bioactive pollutants in water samples, most of the existing bioassays are either very specific for one or a few compounds or are non-specific indicators for general toxic effects.
The model organism
The invertebrate Caenorhabditis elegans has been used as a model organism to predict the toxicity of chemical substances. This soil-dwelling nematode provides particular experimental advantages such as small size, ease to handle, bacterivore, short life cycle, and relatively cheap to maintain in an ordinary laboratory setting. Importantly, it provides the opportunity to study effects of toxicants which alter gene expression (also known as transcriptomics). This is because the nematode’s genome has already been fully sequenced. Additionally, many genes or signaling pathways are conserved between C. elegans and higher organisms, making it suitable as a model for risk assessments.
Sensing contaminants
In this study, the researchers investigated the genetic responses of C. elegans to genotoxic model compounds, including N-ethyl-N-nitrosourea (ENU), formaldehyde (HCHO), and methyl methanesulfonate (MMS) dissolved in water. The study successfully demonstrated that the nematode exposure to compounds resulted in the gene expression profiles that are unique for each toxicant. Among the differentially expressed genes, many were related to the toxicity mechanisms of the tested compounds. Hence, the transcripts of such genes can be used as biomarkers to identify and quantify the toxic potency of single toxicants or mixtures. It was also showed that a relatively high number of genes were downregulated in nematodes treated with toxicants, suggesting a general stress due to the high concentration (5 mM) used for exposure. Surprisingly, no change was found in expression of DNA damage response (DDR) genes. This finding suggests that very high exposure concentrations most likely induce general stress that can mask specific effects of toxicants.
Overall, the study showed that gene expression profiling of C. elegans can provide insights into the type of toxic mechanisms involved and can be translated towards the nature of the toxicants present. The response magnitude can also be related to the exposure concentration.
These results are of interest to researchers since this study can serve as a basis for developing transcriptional biomarkers for detecting a wide array of bioactive contaminants, including hydrophilic ones that are hard to detect chemically. Such an assay can be used as an early warning system that can indicate safety or detect the presence of hazardous chemicals, in e.g. ground-, surface- and potable water.
The paper entitled “Development of a transcription-based bioanalytical tool to quantify the toxic potencies of hydrophilic compounds in water using the nematode Caenorhabditis elegans” was authored by Antoine Karengera, Cong Bao, Joost A.G. Riksen, H. Pieter J. van Veelen, Mark G. Sterken, Jan E. Kammenga, Albertinka J. Murk, Inez J.T. Dinkla and was published in Ecotoxicology and Environmental Safety.